Chemical Shift Imaging (CSI) data interpreter that comes with support for data visualization, spectral processing, spectral localization and estimation, spectral quantification, and multi-variate analytical procedures, such as PCA and cNMF. #Analyze CSI data #CSI data analysis #Interpret CSI data #Analysis #Analyzer #Analyze
3DiCSI is an advanced Windows application developed for helping you examine and interpret Chemical Shift Imaging (CSI) data. It is able to process various multi-dimension data sets, like 1D, 2D single/multi slice, and 3D CSI.
The tool works with different image types, namely Philips (PAR/REC), GE Genesis (pre 11), GE Excite 11.0 or later (DICOM), Siemens (MAG file), Standard DICOM file, and Bruker (2dseq file). You are allowed to import any unknown data type by manually entering the dedicated parameters, and extend the list with file formats with the aid of third-party plugins.
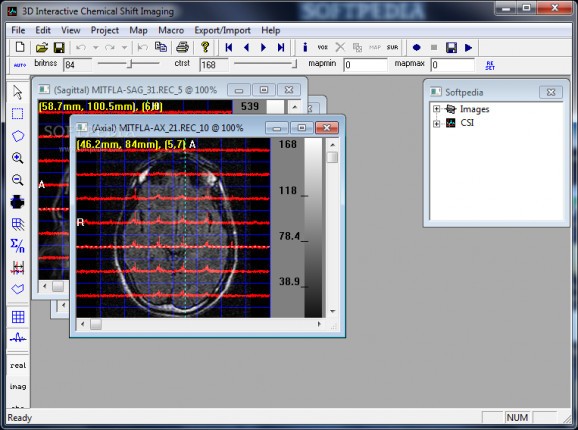
You can check out CSI data and anatomic reference images in the same window, while each voxel can be previewed in a separate panel with its spatial location displayed in three orthogonal image windows.
3DiCSI offers support for smart processing tools which enable you to manipulate data set so it can be phased, zero-filled, spatially smoothed, filtered, and baseline-corrected. The adjustments are applied in real time. The most important utilities worth being mentioned are specialized in constant phase and linear phase correction, temporal filtering (Lorentzian, Gaussian, or combined), spatial smoothing (Hamming, Cosine, Fermi), baseline correction, water suppression, PPM origin adjustment and zero filling, lipid suppression, and peak alignment.
Batch actions can be employed for recording the sequence of processing steps to macro files which can be later on used on CSI data set. Plus, you may apply macro actions to a series of exams in a batch mode.
Project reports can be exported to HTML file format and include information about the image, spectra, metabolic map, and other corresponding parameters. You may print data and save a project to a file on your computer so you can import it in the future.
3DiCSI lets you make use of various analysis methods, such as Principal Component Analysis (PCA) for user-defined spectral region and ROI in order to detect Principal Components (PCs) for the variation sources contained in the CSI data set, PCA-based iterative peak align procedure to get rid of frequency and phase variations, as well as Constrained Non-Negative Matrix Factorization (cNMF) method to recuperate spectral patterns and their spatial distributions.
All in all, 3DiCSI comes packed with a comprehensive suite of features that range from data visualization, spectral processing, spectral localization and estimation, and spectral quantification up to analytical procedures, such as Principal Component Analysis (PCA) and constrained Non-negative Matrix Factorization (cNMF). The advanced package of functions makes it suitable especially for professional users.
3DiCSI 1.9.11
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 3.3 MB
- filename:
- 3DiCSI-inst.exe
- main category:
- Science / CAD
- developer:
- visit homepage
Windows Sandbox Launcher
4k Video Downloader
7-Zip
Bitdefender Antivirus Free
calibre
ShareX
Zoom Client
Microsoft Teams
Context Menu Manager
IrfanView
- Microsoft Teams
- Context Menu Manager
- IrfanView
- Windows Sandbox Launcher
- 4k Video Downloader
- 7-Zip
- Bitdefender Antivirus Free
- calibre
- ShareX
- Zoom Client