A lightweight yet advanced command line application developed to serve in multiple alignment of nucleic acid sequence operations. #Multiple alignment of nucleic acid #Align protein sequences #Protein sequences alignment #View #Analyze #Align
ClustalW is a complex and reliable piece of software developed to provide genetics professionals with an effective method of performing multiple alignment tasks, also being able to create phylogenetic trees.
The application does not benefit from a GUI, which makes it rather unapproachable for inexperienced individuals, since it requires at least some basic knowledge in working in CMD.
Nonetheless, ClustalW comes with a hefty help documentation which can easily be displayed for you to read, on condition that you input the corresponding command.
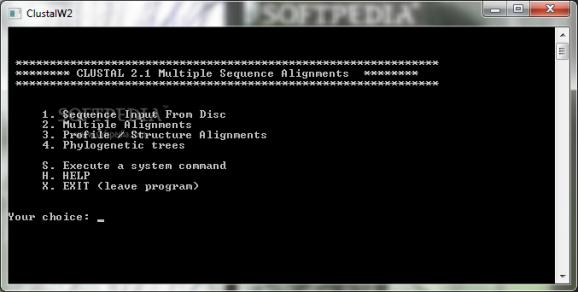
The utility offers several main functions, enabling you to ‘Sequence Input from Disc’, execute ‘Multiple Alignments’ or ‘Profile / Structure Alignments’, as well as generate ‘Phylogenetic Trees’, each operation being assigned a number (1 through 4) which serves as argument.
ClustalW supports a wide array of sequence files, including NBRF-PIR, Fasta, ALN (Clustal), Pileup or GDE, automatically recognizing their format in most of the cases, based on information found at the beginning or end of the document, which is specific to each one.
To add the files you wish to work with, first you need to type ‘1’, then enter the name of the targeted item, along with its full path. Next, you can choose the preferred operation and press the equivalent command, namely ‘2’ for aligning a set of sequences or ‘3’ for inserting an additional sequence into an existing file; command ‘4’ is meant to generate phylogenetic trees from previously aligned sequences.
To sum it up, ClustalW is a useful and quite efficient program, functioning in command line mode to assist you in aligning several different DNA sequences into a single document, with almost no effort for you.
ClustalW 2.1
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 435 KB
- filename:
- clustalw-2.1-win.msi
- main category:
- Science / CAD
- developer:
- visit homepage
7-Zip
4k Video Downloader
IrfanView
Bitdefender Antivirus Free
Zoom Client
Context Menu Manager
calibre
ShareX
Windows Sandbox Launcher
Microsoft Teams
- ShareX
- Windows Sandbox Launcher
- Microsoft Teams
- 7-Zip
- 4k Video Downloader
- IrfanView
- Bitdefender Antivirus Free
- Zoom Client
- Context Menu Manager
- calibre