Flexible, Java-based program with scientific applications in the area of haplotype analysis, with support for various tasks in connection to such process. #Haplotype analysis #Haplotype block analysis #Haplotype association test #Analyze #Analysis #Test
Haplotype analysis is designed for molecular genetic testing in order to identify DNA segments that share similarities. Haploview has been created to improve the tasks relating to such analysis.
The application is Java-based, which makes it cross-platform and there is no need for going through an installation process.
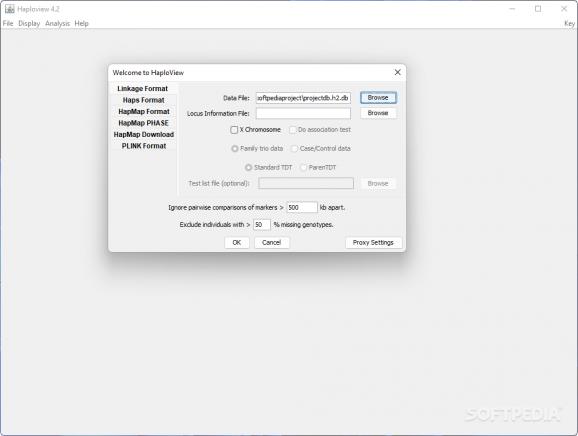
Looks are simple and to the point, with a single panel that accepts the loading of up to five file formats: standard linkage, data dumps from HapMap Project, PHASE or PLINK. There is also support loading multiple files by applying the “-batch” command line.
Regardless of the chosen file format loaded in the application the same parameters can be customized. These refer to the pairwise comparison of markers, which can be above a user specified value (the default is set to 500KB).
A second parameter refers to the amount of genotypes missing in individuals. Depending on the user-set value, these individuals can be excluded from the analysis.
Additional configuration settings are available for some of the supported file formats. In the case of standard linkage users have the possibility to enable association testing, family trio data or load up a test file.
After loading such a file there are various displays available, which include LD plot, haplotypes, check markers or tagger (combines pairwise methods with the potential efficiency of multimarker approaches).
With PHASE format the HapMap info track can be downloaded if there is an Internet connection.
The program can be used for running LD and haplotype analysis as well as estimate the haplotype frequency in population. The list of functions also supports permutation testing that could reveal association significance.
Haploview is a program with scientific applications that supports several tasks relating to the process of haplotype analysis. It can be used by students familiar with the subject and it is also flexible in terms of functionality and usability.
Haploview 4.2
add to watchlist add to download basket send us an update REPORT- PRICE: Free
- runs on:
- Windows All
- file size:
- 5.1 MB
- filename:
- Haploview.jar
- main category:
- Science / CAD
- developer:
Microsoft Teams
ShareX
IrfanView
Windows Sandbox Launcher
Zoom Client
calibre
paint.net
4k Video Downloader
7-Zip
Bitdefender Antivirus Free
- 4k Video Downloader
- 7-Zip
- Bitdefender Antivirus Free
- Microsoft Teams
- ShareX
- IrfanView
- Windows Sandbox Launcher
- Zoom Client
- calibre
- paint.net