An alternative to Modeltest application with support for 24 models of nucleotide substitution that can be implemented in MrBayes tool for estimation of phylogeny. #DNA model #DNA sequence #Substitution model #DNA #Sequence #Nucleotide
Comparing different models of DNA evolution is a task reserved for a particular set of users, who have the necessary knowledge to extract useful information.
They also need specific tools to conduct the operation and MrModeltest is one utility specifically designed for the job.
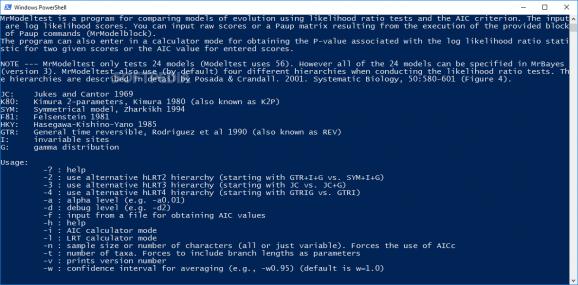
The application does not need to be installed and is console-based. This might distance some of the user from it, although once launched, it presents the proper instructions on using it and the commands and arguments supported.
The developer informs that the tool employs likelihood ratio tests and AIC (Akaike information criterion), which measures a relative quality of a statistical model for specific data.
Additional details provided by the developer refer to the amount of models of nucleotide substitution supported by the application, which is set to 24, and all of them can be specified in MrBayes tool (used for the Bayesian estimation of phylogeny).
Despite a graphical user interface, working with MrModeltest may not be too difficult for the users it targets, as the console window shows the list of supported commands.
The program permits selecting the hLRT2/3/4 hierarchy as well as defining the alpha and debug levels. Examples are presented on the screen to make things easier.
There is also the possibility to enable the AIC or the LRT calculator mode and setting the number of taxa. For most of the operations there are additional tooltips that can help the user.
MrModeltest is a fork of the Modeltest application that supports almost twice the number of models of nucleotide substitution. But its advantage is that these can be implemented in MrBayes utility for phylogeny estimation.
MrModeltest 2.3
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 409 KB
- filename:
- mrmodeltest2_32bit.exe
- main category:
- Science / CAD
- developer:
- visit homepage
4k Video Downloader
paint.net
Zoom Client
Windows Sandbox Launcher
ShareX
calibre
7-Zip
IrfanView
Bitdefender Antivirus Free
Microsoft Teams
- IrfanView
- Bitdefender Antivirus Free
- Microsoft Teams
- 4k Video Downloader
- paint.net
- Zoom Client
- Windows Sandbox Launcher
- ShareX
- calibre
- 7-Zip