A handy and reliable Java-based application designed for extracting mass spectra and peptide sequence data out of X!Tandem XML files. #X!Tandem extractor #Protein identification #Peptide sequence #X!Tandem #Extractor #Parser
X!Tandem is an open source tool that helps microbiologists and geneticists match tandem mass spectra with peptide sequences, in order to identify oligonucleotides and their chemical structure.
X!Tandem Viewer is a nifty piece of software that can help you open the XML files created by X!Tandem and extract relevant information from them. In order to properly work, the program requires Java installed and running on your computer.
The application provides you with an efficient way of opening XML files created by X!Tandem, so that you can extract detailed information about various peptide sequences and genome oligonucleotides. By doing so, you can find data about certain proteins and identify their properties.
You can export your information with ease, either to Spectra file tables or any similar type. By doing so, you can access the information they contain through other specialized applications that deal with proteomic identification.
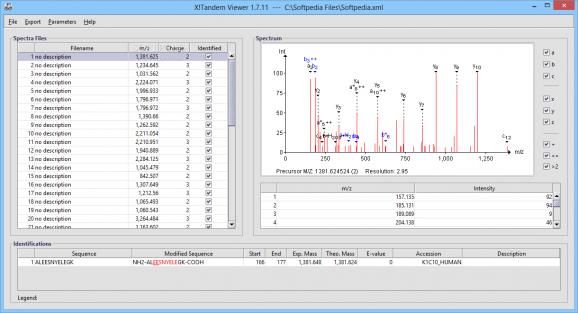
X!Tandem Viewer allows you to access the data stored by X!Tandem XML files and conveniently display it, along with charts and proteic tables. This way, you have the chance to view how certain peptides are related and what is the chemical difference between each one.
Aside from this, the application can show certain proteomic sequences, along with derived peptides and some chemical data, such as atomic mass and accession. In this manner, you can find out extensive information about each scanned oligonucleotide.
To sum it up, X!Tandem Viewer offers you a quick and simple way of reading the contents of the XML files generated by X!Tandem. Displaying both charts and proteomic data, the application will help you better understand the peptide sequences that compose oligonucleotides.
What's new in X!Tandem Viewer 1.13.0:
- LIBRARY UPDATE: Updated utilities to version 4.10.0.
X!Tandem Viewer 1.13.0
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 4.8 MB
- filename:
- xtandem-parser-1.13.0.zip
- main category:
- Science / CAD
- developer:
- visit homepage
4k Video Downloader
Zoom Client
Bitdefender Antivirus Free
ShareX
Context Menu Manager
7-Zip
Microsoft Teams
calibre
Windows Sandbox Launcher
IrfanView
- calibre
- Windows Sandbox Launcher
- IrfanView
- 4k Video Downloader
- Zoom Client
- Bitdefender Antivirus Free
- ShareX
- Context Menu Manager
- 7-Zip
- Microsoft Teams