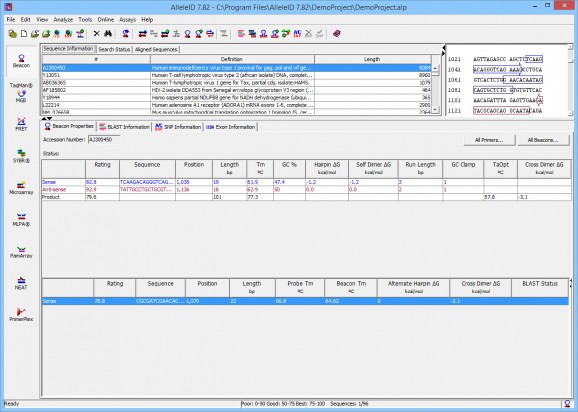
A real time PCR primer and probe design tool that supports bacterial identification, pathogen detection or species identification. #Beacon design #Probe design #Microarray probe #Beacon #Probe #Microarray
Studying bacterias, along with their pathogenic characteristics, takes a considerate amount of time, given that the oligo sequences they contain tend to be pretty long. In order to ease off the time needed to analyze this information, you could use a software tool that has the ability to perform bacterial identification, pathogen detection or species identification processes with ease.
AlleleID is such an application, as it gives you the possibility to analyze assay designs of bacterial oligonucleotides, the determine their pathogenic behavior and species.
The program can help you analyze bacterial oligonucleotide sequences, in order to better understand the behavior or certain bacteria, along with their pathogenic characteristics. You have access to several assays, so that you can investigate the behavior of certain bacterial DNA or RNA molecules from different views, then determine their pathogenic characteristics.
For instance, the program analyzes conserved and species specific regions using one assay then, with the help of certain procedures, such as PCR primer design and dual labeled probe design, detect only the strain or species that interests you from a certain mix.
AlleleID can help you determine the species of a bacteria even if you have a partial oligonucleotide sequence of a gene. This is done by studying the genome draft of related organisms and it allows you to surpass certain difficult tasks, such as detection, identification, quantification or monitoring of contaminants and their environments.
You can identify cross species genomes, the application helping you separate the conserved regions of a oligonucleotide sequence and design an universal probe.
To sum it up, AlleleID confers you with several innovative ways of identifying bacterial genomes along with their pathogenic characteristics and species behavior, using robust assay results and gene splicing analyzers.
What's new in AlleleID 7.85:
- The upgrade accommodates the latest genomic databases available at the NCBI-BLAST server and the recent changes made by NCBI on its web sites and services to improve security and privacy.
AlleleID 7.85
add to watchlist add to download basket send us an update REPORT- runs on:
-
Windows 10 32/64 bit
Windows 8 32/64 bit
Windows 7 32/64 bit
Windows Vista 32/64 bit
Windows XP - file size:
- 108 MB
- filename:
- AL785InstallerWin32.exe
- main category:
- Science / CAD
- developer:
- visit homepage
ShareX
Windows Sandbox Launcher
7-Zip
Context Menu Manager
Zoom Client
Microsoft Teams
Bitdefender Antivirus Free
calibre
IrfanView
4k Video Downloader
- calibre
- IrfanView
- 4k Video Downloader
- ShareX
- Windows Sandbox Launcher
- 7-Zip
- Context Menu Manager
- Zoom Client
- Microsoft Teams
- Bitdefender Antivirus Free