This application features a general purpose multiple sequence alignment program for DNA or proteins, performing comparisons and generating analysis reports. #Multiple sequence alignment #DNA profile alignment #Match sequences #Sequence #Alignment #Profile
Clustal X is an advanced program that deals with multiple sequence alignment for proteins and DNA. Designed as a GUI for ClustalW, the program carries out in-depth sequence analysis, while also offering many utilities to perform proper comparisons.
The program is able to work with ready sequences, imported from files of the following formats: EMBL/SWISSPROT, NBRF/PIR, Pearson (Fasta), GCG/MSF (Pileup), Clustal (*.aln), GCG9/RSF and GDE.
During the import process, Clustal X attempts to detect whether the sequences are nucleotides or amino acids, but there’s no actual guarantee that the choice will be correct.
Once the source file has been loaded, the sequence alignment will be displayed in the main window of the program. The built-in coloring scheme serves to highlight conserved features, making them pop out via vivid colors.
In order to distinguish the contents of the sequences, it is recommended that you use a larger font; this will result in a zoomed sequence view, making it easier to spot its composition.
The operations allowed by the program make quite the list; you can append a sequence to an existing alignment, cut or paste a fragment, as well as to search for a string or to remove existing gaps.
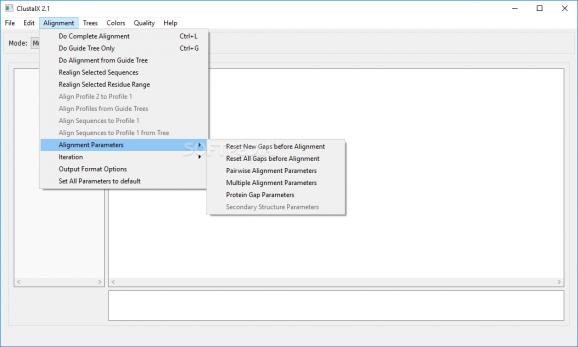
Also, you can perform alignments via one of the following methods: complete (the sequence will undergo a full analysis), guide tree only or a realignment of the selected sequences and / or residue ranges.
It is recommended that before starting an alignment process, you reset all the parameters (new gaps, protein gaps) in order for the results to be conclusive.
One of the most important indicative of the analysis will be the alignment quality score, which is plotted for each column individually. A high score designates a well-conserved column, while low conservation levels are indicated by a lower score.
However, the analysis depends on a variety of parameters, which should be set in accordance with the scientific purpose that needs to be carried out. Most definitely, the program makes a great choice for scientists, as well as students who are looking to make a carrier out of DNA sequencing.
Clustal X 2.1
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 4.7 MB
- filename:
- clustalx-2.1-win.msi
- main category:
- Science / CAD
- developer:
- visit homepage
Zoom Client
Context Menu Manager
Bitdefender Antivirus Free
Microsoft Teams
7-Zip
4k Video Downloader
calibre
ShareX
Windows Sandbox Launcher
IrfanView
- ShareX
- Windows Sandbox Launcher
- IrfanView
- Zoom Client
- Context Menu Manager
- Bitdefender Antivirus Free
- Microsoft Teams
- 7-Zip
- 4k Video Downloader
- calibre