Manage format tree files that feature phylogenetic data supported by MrBayes, Phylip or TreeBase using this simple and straightforward app. #Display phylogenetic tree #View cladogram #Phylogram viewer #Tree file #Phylogenetic tree #Phylogram
The evolution and the relationship between various species can be easily represented through phylogenetic trees, which are also called evolutionary trees.
The information they hold is contained in files of specific format that can be loaded in certain software applications.
TreeView X is a program specifically designed to load up phylogenetic trees. Even if you are not familiar to the subject this digital solution may grab your attention once you manage to load one of the supported file formats (nexus tree file, TreeBase, MrBayes consensus, Clustalx and Phylip).
Installing the application is a simple task that can be easily completed by any user without difficulty.
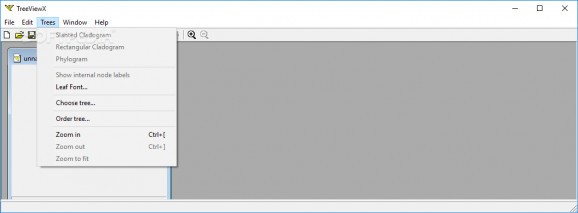
The main screen lacks complexity and makes available all the functions within plain view. All there is needed to benefit from them is a tree file to load up. Finding a sample is not exactly an easy task, despite the affluence of information provided by the Internet these days.
However, once you manage to find the right file TreeView X should display the tree and allow users to easily move from one tree to another and check the slanted or rectangular cladogram, which is one way to see the genealogy between taxa.
The application can also display the internal node labels and provides the possibility to zoom in and out of the image.
Working with it is a simple task that allows different views of the data, such as cascade or tiling it vertically or horizontally.
On the downside, keep in mind that TreeView X is not actively supported by the developer and any issues that may occur from using it may not be fixed.
What's new in TreeView X 0.5:
- Support for picture files (SVG and EMF).
- Some code tidying up.
TreeView X 0.5
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 2.3 MB
- filename:
- setup.exe
- main category:
- Science / CAD
- developer:
7-Zip
ShareX
IrfanView
4k Video Downloader
Microsoft Teams
Context Menu Manager
Windows Sandbox Launcher
Zoom Client
calibre
Bitdefender Antivirus Free
- Zoom Client
- calibre
- Bitdefender Antivirus Free
- 7-Zip
- ShareX
- IrfanView
- 4k Video Downloader
- Microsoft Teams
- Context Menu Manager
- Windows Sandbox Launcher