A Java-enabled application that can help scientists to analyze nucleotide substitutions, with up to five model selection strategies. #Nucleotide substitution #Nucleotide model selection #Likelihood calculator #Nucleotide #Substitution #Analysis
jModelTest is a scientific application that can perform a selection of nucleotide substitutions, useful in DNA analysis, with support for large packets of data.
It is an open-source, cross-platform project that targets students, teachers, as well as researchers in the DNA field. Most of the analysis is done automatically, however, advanced nucleotide knowledge is expected from the user, mostly for the interpretation of the results.
jModelTest relies on the Java Runtime Environment to run successfully, but that’s the only dependency you need to provide. It comes wrapped up inside a portable package, therefore installation is not required.
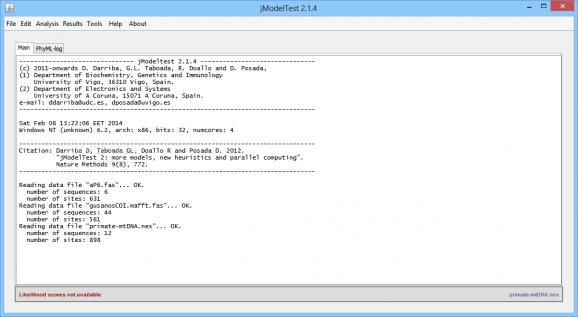
The interface of jModelTest is fairly simple, but the functions embedded inside it are of a more complex nature. Most of the GUI is dedicated to the analysis of the data and the reports that result from this process, which, depending on the information contained inside the source file, can be quite lengthy.
There are five different selection strategies and these are embedded inside the Analysis menu, as follows: AIC (Akaike Information Criterion), BIC (Bayesian Information Criterion), DT (Decision Theory), hLRT (hierarchical likelihood ratio tests) and Phylogenic Averaging.
All of these can be configured before the actual analysis process; for instance, you can adjust the confidence interval or set the application to perform a model averaging. To top it off, you can also instruct the program to do a likelihood score computation that depends on a customizable rate variation and a pre-selected base tree model.
In conclusion, jModelTest can save the time and effort scientists would invest in performing DNA analysis operations by traditional means. However, it is recommended that the results be thoroughly verified, just in case.
What's new in jModelTest 2.1.7:
- Fixed bug with special characters in paths
- Added initial check of PhyML binaries
- Added notification in case AICc produces negative values
- Allow for environment variables on configuration file
jModelTest 2.1.7
add to watchlist add to download basket send us an update REPORT- runs on:
- Windows All
- file size:
- 10.1 MB
- filename:
- jmodeltest-2.1.7.tar.gz
- main category:
- Science / CAD
- developer:
- visit homepage
Bitdefender Antivirus Free
Microsoft Teams
7-Zip
calibre
Zoom Client
IrfanView
4k Video Downloader
Context Menu Manager
Windows Sandbox Launcher
ShareX
- Context Menu Manager
- Windows Sandbox Launcher
- ShareX
- Bitdefender Antivirus Free
- Microsoft Teams
- 7-Zip
- calibre
- Zoom Client
- IrfanView
- 4k Video Downloader